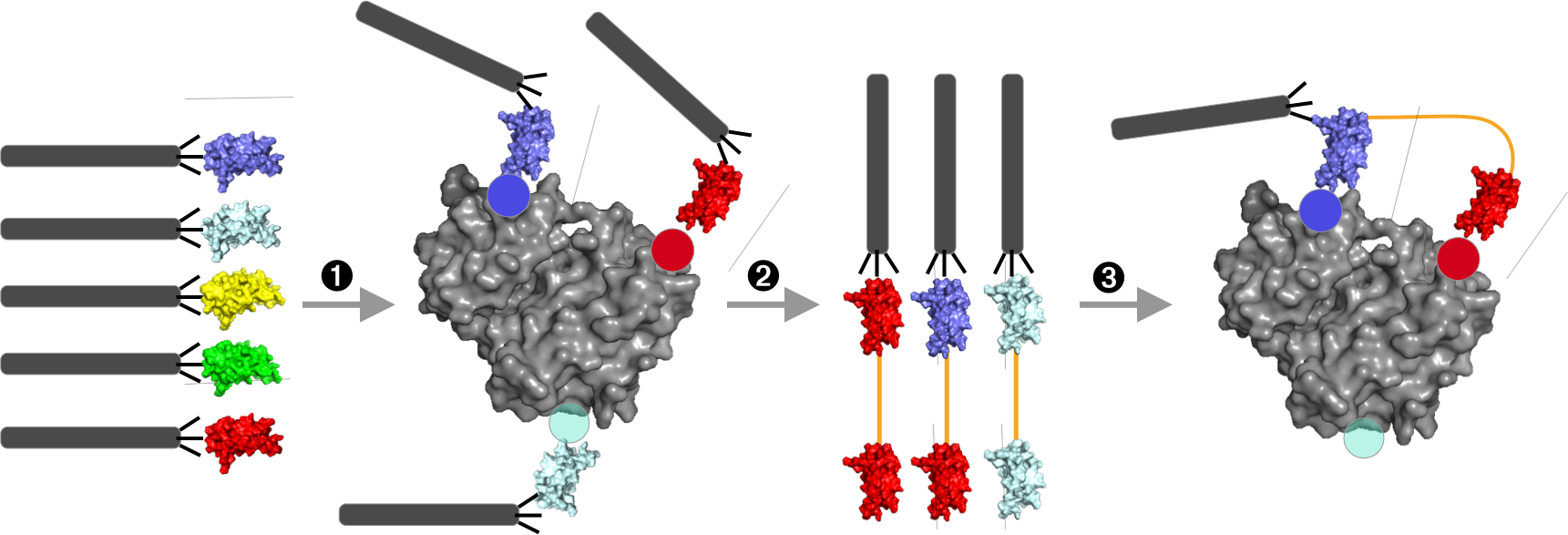
Fig. 1 – MegaSTAR process
Step 1: Affinity selection with the monovalent phage-display library generates a pool of potential binders (black filamentous virions displaying blue, cyan, and red monobodies). Different colored monobodies refer to different sequence variants that bind different epitopes on the target (gray). Step 2: The output clone pool is converted into randomly-combined pairs of monobodies linked through a flexible glycine-serine linker (orange). Step 3: Through additional rounds of affinity selection with low amounts of targets, tandem pairs that bind to two non-overlapping epitopes will predominate, as their affinity will be dramatically increased through avidity (i.e., simultaneous binding to two epitopes).
Tango Biosciences uses proprietary technology for discovering pairs of recombinant affinity reagents through phage-display. This process (Fig. 1) is termed Megaprimer Shuffled Tandem Affinity Reagents (MegaSTAR). MegaSTAR is innovative because it identifies between 5 and 15 two-site capable binders (i.e., pairs) as part of the affinity selection process in phage-display. By relying upon two independent affinity reagents that recognize distinct epitopes, the sandwich assay increases both sensitivity and specificity.
Millions of pair-wise combinations of monobodies are created during MegaSTAR, and stringent affinity selections enrich for two-side capable reagents. Not only does this save time and money, but more importantly we discover many more pairs of two-site capable reagents than through traditional phage-display, which biases towards high affinity binders at the exclusion of moderate to weak binders of novel epitopes. One successful example is Pak1, in which we have generated 8 unique pairs of binders at 7 different epitopes.
Gorman KT, Roby LC, Giuffre A, Huang R, Kay BK. Tandem phage-display for the identification of non-overlapping binding pairs of recombinant affinity reagents. Nucleic Acids Res. 2017; 45(18) e158. doi:10.1093/nar/gkx688. PubMed PMID: 28985360.
